skg.ngauss¶
N-dimensional, unnormalized Gaussian fit.
This method is adapded from the references to apply to any number of dimensions, and arbitrary orientation of the error ellipsoid.
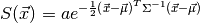
Here, 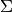 is the covariance matrix that describes the
ellipsoid of the Gaussian. If this were a density function, the
amplitude
is the covariance matrix that describes the
ellipsoid of the Gaussian. If this were a density function, the
amplitude a would be fixed at
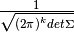 .
.
This function is specially suited for work with images. A wrapper to
facilitate image inputs is therefore provided:
ngauss_from_image.
Functions
anthony_weights([noise]) |
Returns a function that computes weights for ngauss_fit based on the noise standard deviation. |
model(x, a, mu, sigma[, axis]) |
Compute 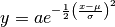 . . |
ngauss_fit(x, y[, axis, weights, scaling]) |
An N-dimensional Gaussian fit. |
ngauss_from_image(img[, weights, scaling]) |
Compute a Gaussian fit to an entire image. |
-
skg.ngauss.anthony_weights(noise=None)[source]¶ Returns a function that computes weights for
ngauss_fitbased on the noise standard deviation.Weights are computed (with appropriate clipping) according to
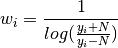
Signals smaller than noise are discarded by setting the weight to zero.
Parameters: noise (float, optional) – An estimate of the standard deviation of signal noise. If not specified, use the standard deviation of y. Returns: weights – A function that accapts x and y and returns an array of weights the same size as y. x is completely ignored. This function is suitable as a weights parameter to ngauss_fit.Return type: callable References
-
skg.ngauss.model(x, a, mu, sigma, axis=-1)[source]¶ Compute
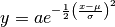 .
.The number of dimensions, N, is determined from
x.shape[axis]. mu must be a vector of length N, and sigma must be an NxN positive-definite matrix.Parameters: - x (array-like) – The value of the model will be the same shape as the input, with axis reduced.
- a (float) – The amplitude at
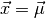 .
. - mu (array-like) – The location of the peak. Must be a an array of length
x.shape[axis]. May be a scalar if the location is the same value in all dimensions. This feature should only be used for a peak at zero. - sigma (float) – The covariance matrix of the Gaussian. Must be a
positive-definite matrix of shape
(x.shape[axis], x.shape[axis]). - axis (int) – The axis corresponding to the dimension of
xthat contains the point vectors.
Returns: y – An array of the same shape as x with axis reduced, containing the model computed for the given parameters.
Return type: array-like
Example
To generate a 100px x 100px 2D image with a spot in the middle:
>>> import numpy as np >>> from skg import ngauss_fit >>> x = np.indices((100, 100), dtype=float) >>> cov = np.array([[100., 40.], [40., 64.]]) >>> img = ngauss_fit.model(x, 255, (50, 50), cov, axis=0)
-
skg.ngauss.ngauss_fit(x, y, axis=-1, weights=None, scaling=False)[source]¶ An N-dimensional Gaussian fit.
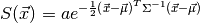
This implementation is based on an extension of [AnthonyGranick] and the first of two passes described in [Wan-Wang-Wei-Li-Zhang].
A weighing function must be applied to the data to avoid having the low-SNR data dominate the fit. The default is to weight the measurements by their intensity, as per _[Wan-Wang-Wei-Li-Zhang]. However, other schemes are possible, such as the one proposed by _[Anthony-Granick]. The latter scheme can be used by passing in a callable returned by
anthony_weightsas the weights parameter.Parameters: - x (array-like) – The x-values of the data points. The fit will be performed on a version of the array raveled across all dimensions except axis.
- y (array-like) – The y-values of the data points corresponding to x. Must be the same size as x except at axis. The fit will be performed on a raveled version of this array.
- axis (int) – The axis containing the vectors of x. The dimension of the
Gaussian is
x.shape[axis]. The default is-1. - weights (array-like or callable, optional) – Either an array with the same number of elements as y (it will be raveled), or a callable that accepts reshaped versions of x and y and returns an array of weights.
- scaling (bool, optional) – If True, scale and offset the data to a bounding box of -1 to +1 in each axis during computations for numerical stability. Default is False.
Returns: - a (float) – The amplitude of the Gaussian.
- mu (~numpy.ndarray) – The mean of the Gaussian, as an N-element array.
- sigma (~numpy.ndarray) – The covariance of the Gaussian, as an NxN positive definite matrix.
See also
ngauss_from_image- A wrapper for fitting over an entire array.
anthony_weights- A function that returns an alternative, noise-specific weighting scheme.
Notes
Negative and zero weights are discarded from the computation without ever being inserted into the solution matrix.
References
-
skg.ngauss.ngauss_from_image(img, weights=None, scaling=True)[source]¶ Compute a Gaussian fit to an entire image.
Parameters: - img (array-like) – The image to process. Usually a segment of a 2D image. The data is expected to have been background subtracted and thresholded so that any low-SNR pixels are set to zero.
- weights (array-like or callable, optional) – A weighing function must be applied to the data to avoid having the low-SNR data dominate the fit. The default is to weight the measurements by their intensity, as per [Wan-Wang-Wei-Li-Zhang]. However, other schemes are possible, such as the one proposed by [Anthony-Granick]. weights can be passed in as an array with the same number of elements as y (it will be raveled), or a callable that accepts reshaped versions of x and y and returns an array of weights.
- scaling (bool, optional) – If True, scale and offset the data to a bounding box of -1 to +1 in each axis during computations for numerical stability. Default is True.
Returns: - a (float) – The amplitude of the Gaussian.
- mu (~numpy.ndarray) – The mean of the Gaussian, as an N-element array.
- sigma (~numpy.ndarray) – The covariance of the Gaussian, as an NxN positive definite matrix.