skg.ngauss_fit¶
-
skg.ngauss_fit(x, y, axis=-1, weights=None, scaling=False)[source]¶ An N-dimensional Gaussian fit.
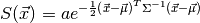
This implementation is based on an extension of [AnthonyGranick] and the first of two passes described in [Wan-Wang-Wei-Li-Zhang].
Parameters: - x (array-like) – The x-values of the data points. The fit will be performed on a version of the array raveled across all dimensions except axis.
- y (array-like) – The y-values of the data points corresponding to x. Must be the same size as x except at axis. The fit will be performed on a raveled version of this array.
- axis (int) – The axis containing the vectors of x. The dimension of the
Gaussian is
x.shape[axis]. The default is-1. - weights (array-like or callable, optional) – A weighing function must be applied to the data to avoid having the low-SNR data dominate the fit. The default is to weight the measurements by their intensity, as per _[Wan-Wang-Wei-Li-Zhang]. However, other schemes are possible, such as the one proposed by _[Anthony-Granick]. weights can be passed in as an array with the same number of elements as y (it will be raveled), or a callable that accepts reshaped versions of x and y and returns an array of weights.
- scaling (bool, optional) – If True, scale and offset the data to a bounding box of -1 to +1 in each axis during computations for numerical stability. Default is False.
Returns: - a (float) – The amplitude of the Gaussian.
- mu (~numpy.ndarray) – The mean of the Gaussian, as an N-element array.
- sigma (~numpy.ndarray) – The covariance of the Gaussian, as an NxN positive definite matrix.
See also
ngauss_from_image- A wrapper for fitting over an entire array.
anthony_weights- A function that returns an alternative, noise-specific weighting scheme.
References
- [Anthony-Granick]
- [Wan-Wang-Wei-Li-Zhang]
- N-Dimensional Generalization of Linearized Gaussian Fitting, Expanding to Arbitrary Dimensions
Notes
Negative and zero weights are discarded from the computation without ever being inserted into the solution matrix.